Coronavirus disease 2019, or COVID-19, is an infectious disease caused by the SARS-CoV-2 virus. It can be very contagious and can spread quickly. Since 2020, more than 1.2 million people have died of COVID-19 in the U.S.
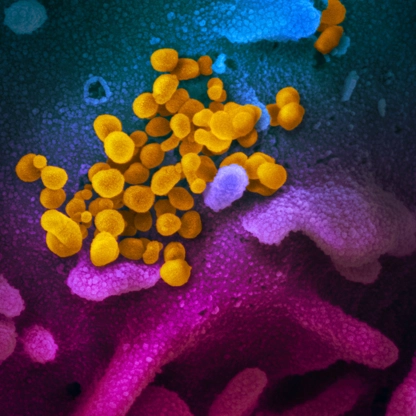
Coronavirus disease 2019, or COVID-19, is an infectious disease caused by the SARS-CoV-2 virus. It can be very contagious and can spread quickly. Since 2020, more than 1.2 million people have died of COVID-19 in the U.S.
People with COVID-19 may have a wide range of symptoms ranging from mild symptoms to severe illness. Possible symptoms include fever or chills, cough, shortness of breath or difficulty breathing, sore throat, congestion or runny nose, new loss of taste or smell, fatigue, muscle or body aches, headache, nausea or vomiting, and diarrhea.
Several antiviral medications are available that can reduce risk of hospitalization and death for people with COVID-19 who are more likely to get very sick. To be effective, treatments must be started within five to seven days after symptoms first develop.
People who are more likely to get very sick include adults over 65, people who are not vaccinated or not up to date on their COVID-19 vaccinations, or people with certain medical conditions, such as chronic lung disease, heart disease or a weakened immune system.
There are several core prevention strategies that everyone can use to help to protect themselves and others from respiratory viruses, including COVID-19. These include good hygiene (covering your coughs and sneezes, washing or sanitizing your hands often, and cleaning frequently touched surfaces), taking steps for cleaner air and staying home when sick.
Staying up to date on COVID-19 vaccines significantly lowers the risk of getting very sick, being hospitalized or dying from COVID-19. The COVID-19 vaccine is recommended for everyone ages 6 months and older and is regularly updated to protect against the currently circulating viral strains. It is especially important to get an updated COVID-19 vaccine if you are aged 65 and older, or if you are at high risk for severe COVID-19. COVID-19 vaccination has also been shown to lower your risk of developing long COVID.
Additional strategies like wearing a mask and putting distance between yourself and others can help lower the risk of spreading COVID-19 and other respiratory viruses. Testing for COVID-19 can help you decide what to do next, like getting treatment to reduce your risk of severe illness and taking steps to lower your chances of spreading COVID-19 to others.
Loading...